Bioinformatics & Computational Biology
We are developing bioinformatics techniques and tools for
uncovering the molecular-level pathways involved in complex diseases such as
cancer, aiming at determining disease markers and therapeutic targets.
- knowledge discovery and machine
learning (especially for extracting functional information from
microarray gene expression data, reconstructing/refining pathways from
large-scale microarray data)
- Biclustering and metaclustering gene expression data
- Genomic subclassification
of cancer (lung, colon, breast, glioblastoma)
- Gene network inference.
- Microarray analysis of cancer
datasets
- pancreatic adenocarcinoma (collaboration with the Fundeni
Clinical Institute) [43]
- colon
cancer [40,44]
- gastric
cancer (with Stefan S. Nicolau
Institute of Virology and Fundeni Clinical Institute) [45]
- lung
cancer (Bhattacharjee and Garber
datasets) [37,32]
- breast
cancer (Pawitan dataset)
- glioblastoma.
- Text mining, intelligent query
answering, ontologies
(mediator architecture providing reasoning-aware query answering)
- Determining potential disease markers
and drug targets from a combination of pathways and gene
expression data.
Bioinformatic analysis of a large pancreatic cancer dataset (GENOPACT project)
Pancreatic ductal adenocarcinoma (PDAC) is one of the deadliest form of cancer, for which the best known therapeutic options
are currently extremely ineffective. Moreover, the precise details of PDAC
pathogenesis are still insufficiently known, requiring the use of
high-throughput methods. In the framework of the GENOPACT
project (Research of Excellence Program CEEX 56/2005), we analyzed a set of
78 pancreatic cancer-normal sample pairs from the tissue bank of the Fundeni Clinical
Institute (ICF), measured with Affymetrix U133 Plus 2.0 microarrays. This is one of the largest available
pancreatic ductal adenocarcinoma
datasets, thereby allowing a statistically reliable identification of the genes
involved in this disease.
We have developed a complex bioinformatic
framework for the analysis of this dataset including:
- a
number of different clustering algorithms, including widely used ones such
as hierarchical clustering, but also original biclustering
algorithms allowing for overlapping clusters
- promoter
analysis for detection of transcription factor binding sites, useful for
determining the regulatory programs of the differentially expressed genes
- a large database of gene interactions and pathways compiled from
several sources, including Pubmed abstracts and
literature.
We have performed an in-depth integrated analysis of the
resulting set of differentially expressed genes, producing a plausible
“model” of the molecular-level mechanisms of PDAC and its
progression.
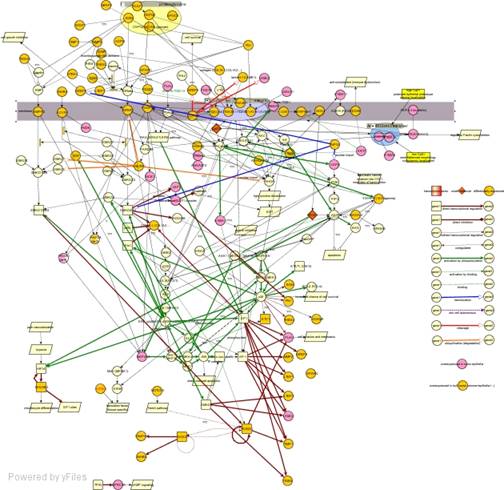
PDAC is especially difficult to study
using microarrays due to its strong desmoplastic
reaction, which involves a hyperproliferating stroma that effectively "masks" the contribution
of the minoritary neoplastic
epithelial cells. Thus it is not clear which of the genes that have been found
differentially expressed between normal and whole tumor tissues are due to the
tumor epithelia and which simply reflect the differences in cellular composition.
To address this problem, laser microdissection
studies have been performed, but these have to deal with much smaller tissue
sample quantities and therefore have significantly higher experimental noise.
We have combined our own large sample
whole-tissue study with a previously published smaller sample microdissection study by Grutzmann
et al. to identify the genes that are specifically overexpressed
in PDAC tumor epithelia.
We have found a number of genes
whose over-expression appears to be inversely correlated with patient survival
[43]:
- keratin
7 (KRT7)
- laminin
gamma 2 (LAMC2)
- stratifin
(SFN)
- annexin
A2 (ANXA2)
- MAP4K4
- platelet
phosphofructokinase (PFKP)
- OACT2 (MBOAT2)
which are all specifically upregulated in the neoplastic
epithelia, rather than the tumor stroma.
We plan to further refine our current
understanding of the molecular-level processes responsible in this disease in
the framework of future projects and with the help of a specialized
molecular-biology lab by using various high-throughput technologies (not just
microarrays) to dissect the pathways involved in PDAC and to test these on cell
lines and possibly animal models.
Bioinformatic analysis of the lung cancer dataset of Bhattacharjee
et al.
-
microarray
data analysis (differentially expressed genes, biclustering,
metaclustering, gene network inference)
-
literature
analysis tools (e.g. extracting co-citations)
The microarray data analysis tools reveal only the level of transcription
regulation and are strongly affected by noise and normal biological
variability. We are therefore using them in conjunction with literature
analysis tools for
-
validating
certain transcriptional influences, as well as
-
emphasizing the various (signaling) pathways in which these genes
operate.
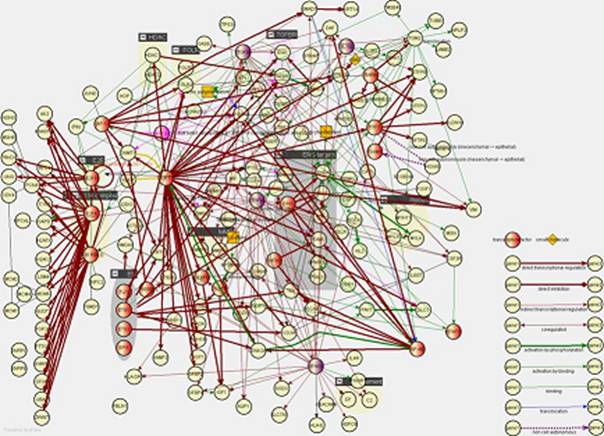
The partial results are very encouraging. For example, for
the squamous cell lung carcinoma we have found
essentially two groups of differentially expressed genes:
-
a
set of upregulated genes involved e.g. in the cell
cycle (e.g. E2F and/or p130/retinoblastoma like 2 targets) and/or in the structure and organization of the
cytoskeleton (e.g. keratin 5, desmoplakin –
specific to the squamous cancer subtype)
-
a larger set of down-regulated genes, normally involved in certain
developmental stages of the lung.
Apparently, this cancer subtype seems to be due to a
defective re-enactment of normal developmental processes (at a wrong time and
place).
References
•
Liviu Badea, Vlad Herlea,
Simona Dima, Traian Dumitrascu, Irinel Popescu. Combined gene expression analysis of whole-tissue and microdissected
pancreatic ductal adenocarcinoma identifies genes specifically overexpressed in
tumor epithelia. Hepatogastroenterology. 2008 Nov-Dec;55(88):2016-27.
•
Liviu Badea. Extracting Gene Expression Profiles Common
to Colon and Pancreatic Adenocarcinoma Using
Simultaneous Nonnegative Matrix Factorization. Proc. Pacific Symposium on Biocomputing PSB-2008, pp. 267-278, World Scientific 2008.
•
Liviu Badea. Generalized Clustergrams for
Overlapping Biclusters. Proceedings of the International Joint Conference on
Artificial Intelligence IJCAI-09, Pasadena, pp 1383-1388, 2009.
•
Liviu Badea. Combining Gene Expression and Transcription
Factor Regulation Data using Simultaneous Nonnegative Matrix Factorization. Proc.
BIOCOMP-2007, pp. 127-131, CSREA Press 2007.
•
Liviu Badea, Doina Tilivea.
Stable
Biclustering of Gene Expression Data with Nonnegative
Matrix Factorizations. Proceedings of
the International Joint Conference on Artificial Intelligence IJCAI-07,
Hyderabad, India, AAAI Press, 2007, pp. 2651-2656. ISSN: 0921-7126
•
Liviu Badea. Combining DNA Copy Number and Gene
Expression Data to Reveal Sample-Specific Genetic Abnormalities in Pancreatic
Cancer. Studies in Informatics and Control, Vol. 15 No. 4, (2006),
pp.403-413, ISSN 1220-1766.
•
Liviu Badea, Doina Tilivea, Meta-clustering
Gene Expression Data with Positive Tensor Factorizations. Proceedings
European Conference on Artificial Intelligence ECAI-06, p. 787, IOS Press 2006.
•
Liviu Badea, Semantic Web reasoning for analyzing gene expression
profiles. PRINCIPLES AND PRACTICE OF
SEMANTIC WEB REASONING. LECTURE NOTES IN COMPUTER SCIENCE, Vol. 4187, pp.
78-89, Springer 2006.
•
Liviu Badea, Doina Tilivea.
Sparse Factorizations of Gene Expression Data guided
by Binding Data. Proceedings of the Pacific Symposium on Biocomputing
PSB-2005, pp. 447-458.
•
F. Bry, C. Koch, T. Furche, S. Schaffert, L. Badea, S. Berger: Querying the Web Reconsidered: Design
Principles for Versatile Web Query Languages. INTERNATIONAL JOURNAL ON
SEMANTIC WEB AND INFORMATION SYSTEMS 1(2):
pp. 1-21 (2005) ISSN: 1552-6283
•
Liviu Badea, Clustering and metaclustering
with nonnegative matrix decompositions.
MACHINE LEARNING: ECML 2005, PROCEEDINGS. LECTURE NOTES IN ARTIFICIAL
INTELLIGENCE, 3720, pp. 10-20 Springer, 2005.
•
Liviu Badea, Determining the Direction of Causal
Influence in Large Probabilistic Networks: A Constraint-Based Approach.
Proceedings of the 14th European Conference on Artificial Intelligence ECAI
2004, IOS Press, pp. 263-267.
•
Badea, L, Tilivea, D, Hotaran, A, Semantic Web
reasoning for ontology-based integration of resources. PRINCIPLES AND
PRACTICE OF SEMANTIC WEB REASONING, PROCEEDINGS. LECTURE NOTES IN COMPUTER
SCIENCE, Vol. 3208, pp. 61-75, Springer, 2004.
•
Rolf Backofen, Mike Badea, Pedro Barahona, Liviu Badea, François Bry, Gihan Dawelbait, Andreas Doms, François Fages, Carole
Goble, Andreas Henschel, Anca
Hotaran, Bingding Huang,
Ludwig Krippahl, Patrick Lambrix,
Werner Nutt, Michael Schroeder, Sylvain Soliman,
Sebastian Will. Towards a semantic web
for bioinformatics. (Poster) In: Proceedings of "Bioinformatics
2004", Linköping, Sweden (3rd - 6th June 2004), SocBIN - Society for Bioinformatics in the Nordic
countries.
•
Liviu Badea. Functional Discrimination of Gene Expression Patterns
in Terms of the Gene Ontology. Pacific
Symposium on Biocomputing 2003: 565-576
•
Liviu Badea, Doina Tilivea, Integrating
biological process modelling with gene expression
data and ontologies for functional genomics (position
paper). COMPUTATIONAL METHODS IN SYSTEMS BIOLOGY, PROCEEDINGS. LECTURE
NOTES IN COMPUTER SCIENCE, Vol. 2602, pp. 187-193, Springer Verlag
2003.
•
Liviu Badea, Doina Tilivea, Intelligent
Information Integration as a Constraint Handling Problem Proc. of the Fifth
International Conference on Flexible Query Answering Systems (FQAS-2002),
Lecture Notes In Computer Science, Vol. 2522, pp. 12-27, Springer Verlag, 2002. ISBN:3-540-00074-7.